-Search query
-Search result
Showing 1 - 50 of 7,826 items for (author: wan & r)

EMDB-39126: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
Method: single particle / : Lin SC, Yang CY

EMDB-39127: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
Method: single particle / : Lin SC, Yang CY

PDB-8ybx: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
Method: single particle / : Lin SC, Yang CY

EMDB-19798: 
human PLD3 homodimer structure
Method: single particle / : Lammens K

EMDB-43753: 
Yeast U1 snRNP with humanized U1C Zinc-Finger domain
Method: single particle / : Shi SS, Kuang ZL, Zhao R

PDB-8w2o: 
Yeast U1 snRNP with humanized U1C Zinc-Finger domain
Method: single particle / : Shi SS, Kuang ZL, Zhao R

EMDB-38215: 
Human GPR34 -Gi complex bound to S3E-LysoPS
Method: single particle / : Kawahara R, Shihoya W, Nureki O

EMDB-38217: 
Human GPR34 -Gi complex bound to S3E-LysoPS, receptor focused
Method: single particle / : Kawahara R, Shihoya W, Nureki O

PDB-8xbe: 
Human GPR34 -Gi complex bound to S3E-LysoPS
Method: single particle / : Kawahara R, Shihoya W, Nureki O

PDB-8xbg: 
Human GPR34 -Gi complex bound to S3E-LysoPS, receptor focused
Method: single particle / : Kawahara R, Shihoya W, Nureki O

EMDB-28941: 
HIV Env BG505_MD39_B11 SOSIP boosting trimer in complex with B11_d77.7 mouse Fab and RM20A3 Fab
Method: single particle / : Torres JL, Ozorowski G, Ward AB

EMDB-28942: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized BG18HCgl knock-in mice
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-28945: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized wild type mice, and NHP monoclonal Fab RM20A3
Method: single particle / : Ozorowski G, Ward AB

EMDB-43190: 
HIV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-44644: 
SARS CoV2 spike in complex with NTD-directed antibody Fab DH1052
Method: single particle / : Edwards RJ, Mansouri K

EMDB-43488: 
Cryo-EM structure of human invariant chain in complex with HLA-DR15
Method: single particle / : Wang N, Caveney NA, Jude KM, Garcia KC

EMDB-43501: 
Cryo-EM structure of human invariant chain in complex with HLA-DQ
Method: single particle / : Wang N, Caveney NA, Jude KM, Garcia KC

PDB-8vrw: 
Cryo-EM structure of human invariant chain in complex with HLA-DR15
Method: single particle / : Wang N, Caveney NA, Jude KM, Garcia KC

PDB-8vsp: 
Cryo-EM structure of human invariant chain in complex with HLA-DQ
Method: single particle / : Wang N, Caveney NA, Jude KM, Garcia KC

EMDB-41639: 
Langya henipavirus fusion protein in postfusion state
Method: single particle / : Wang Z, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-41640: 
Langya henipavirus fusion protein in prefusion state
Method: single particle / : Wang Z, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-41641: 
Langya henipavirus postfusion F protein in complex with the 4G5 Fab, local refinement of the viral membrane distal region
Method: single particle / : Wang Z, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-41642: 
Langya henipavirus postfusion F protein in complex with 4G5 Fab, local refinement of the viral membrane proximal region
Method: single particle / : Wang Z, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-41644: 
Langya henipavirus postfusion fusion protein in complex with 4G5 Fab (global refinement)
Method: single particle / : Wang Z, Veesler D

PDB-8tve: 
Langya henipavirus fusion protein in postfusion state
Method: single particle / : Wang Z, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8tvf: 
Langya henipavirus fusion protein in prefusion state
Method: single particle / : Wang Z, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8tvg: 
Langya henipavirus postfusion F protein in complex with the 4G5 Fab, local refinement of the viral membrane distal region
Method: single particle / : Wang Z, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-8tvh: 
Langya henipavirus postfusion F protein in complex with 4G5 Fab, local refinement of the viral membrane proximal region
Method: single particle / : Wang Z, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-40861: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE
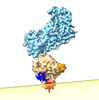
EMDB-40901: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the PH domain
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE
EMDB-40902: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the second PH domain
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-40903: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state (full helix)
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-40942: 
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-40946: 
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state (full helix)
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

PDB-8sxz: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

PDB-8sz4: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the PH domain
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

PDB-8sz7: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the second PH domain
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

PDB-8sz8: 
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state (full helix)
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

PDB-8t0k: 
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

PDB-8t0r: 
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state (full helix)
Method: helical / : Jimah JR, Canagarajah BJ, Hinshaw JE

EMDB-34992: 
Cryo-EM Structure of CdnG-E2 complex from Serratia marcescens (UltrAuFoil)
Method: single particle / : Xiao J, Wang L

EMDB-39353: 
Cryo-EM Structure of CdnG-E2 complex from Serratia marcescens
Method: single particle / : Xiao J, Wang L

PDB-8hsb: 
Cryo-EM Structure of CdnG-E2 complex from Serratia marcescens (UltrAuFoil)
Method: single particle / : Xiao J, Wang L

PDB-8yjy: 
Cryo-EM Structure of CdnG-E2 complex from Serratia marcescens
Method: single particle / : Xiao J, Wang L

EMDB-37963: 
Cryo-EM structure of the hamster prion 23-144 fibril at pH 3.7
Method: helical / : Lee CH, Saw JE, Chen E, Wang CH, Chen R

PDB-8wzx: 
Cryo-EM structure of the hamster prion 23-144 fibril at pH 3.7
Method: helical / : Lee CH, Saw JE, Chen E, Wang CH, Chen R

EMDB-37342: 
Structural mechanism of inhibition of the Rho transcription termination factor by Rof
Method: single particle / : Zhang J, Wang C

PDB-8w8d: 
Structural mechanism of inhibition of the Rho transcription termination factor by Rof
Method: single particle / : Zhang J, Wang C

EMDB-41636: 
Ghanaian virus fusion glycoprotein (GhV F)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
